eugenol
Identification (Source:EPA,EU)
| ID |
1296 |
| Name |
eugenol |
| CAS |
97-53-0 |
| IUPAC |
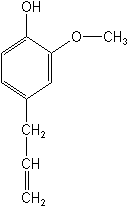
4-allyl-2-methoxyphenol |
| Inchi |
InChI=1S/C10H12O2/c1-3-4-8-5-6-9(11)10(7-8)12-2/h3,5-7,11H,1,4H2,2H3 |
| CIPAC |
|
| EPA Code |
|
| Formula |
C10H12O2 |
| SMILES |
O(C([H])([H])[H])c(c(c([H])c([H])c1C([H])([H])C([H])=C([H])[H])O[H])c1[H] |
| Status |
PENDING |
Classification
Properties (Source:PubChem)
| Molecular Weight |
164.20107 |
| Physical State |
|
| AlogP |
2.579 |
| H-Bond Donor |
1 |
| H-Bond Acceptor |
2 |
| Surface Area |
175.53 |
| Polar Surface Area |
29.46 |
Structure
Related targets
| Target Name |
RScore (About this table) |
|
|
vanilloid receptor 1
|
721(8,10,11,16) |
Details |
|
cyclooxygenase 2
|
709(8,9,11,29) |
Details |
|
xanthine oxidase
|
572(6,8,10,22) |
Details |
|
5 lipoxygenase
|
351(3,6,7,16) |
Details |
|
cytochrome P450 (protein family or complex)
|
305(3,4,6,25) |
Details |
|
monoamine oxidase A
|
275(3,4,4,5) |
Details |
|
IgE
|
255(3,3,4,10) |
Details |
|
aryl hydrocarbon receptor
|
247(3,3,3,7) |
Details |
|
Lipoxygenase (protein family or complex)
|
260(2,5,5,10) |
Details |
|
calcium channel (protein family or complex)
|
225(2,4,4,5) |
Details |
|
TRPV3
|
201(2,3,3,11) |
Details |
|
myeloperoxidase
|
200(2,3,4,5) |
Details |
|
Ca v
|
200(2,3,4,5) |
Details |
|
E2F1
|
194(2,3,3,4) |
Details |
|
cytochrome P450 1A1
|
193(2,3,3,3) |
Details |
|
CYP2D6
|
180(2,2,5,5) |
Details |
|
IL 1beta
|
177(2,2,2,17) |
Details |
|
alkaline phosphatase
|
177(2,2,3,12) |
Details |
|
TNFalpha
|
170(2,2,2,10) |
Details |
|
mitogen activated protein kinases (protein family or complex)
|
170(2,2,3,5) |
Details |
|
NF kappa B
|
162(2,2,2,2) |
Details |
|
ATPase
|
184(1,4,4,14) |
Details |
|
Acetylcholinesterase
|
181(1,4,4,11) |
Details |
|
glutathione S transferase
|
166(1,3,6,11) |
Details |
|
Tyrosinase
|
160(1,3,3,20) |
Details |
|
XL1
|
149(1,3,4,4) |
Details |
|
lipase
|
146(1,3,3,6) |
Details |
|
bcl 2
|
144(1,3,3,4) |
Details |
|
aldose reductase
|
118(1,2,2,8) |
Details |
|
sodium channel (protein family or complex)
|
118(1,2,3,3) |
Details |
|
demethylase
|
114(1,2,2,4) |
Details |
|
H:quinone oxidoreductase
|
113(1,2,2,3) |
Details |
|
cyclooxygenase 1
|
113(1,2,2,3) |
Details |
|
UGT2B15
|
94(1,1,1,14) |
Details |
|
UGT2B17
|
91(1,1,1,11) |
Details |
|
microsomal epoxide hydrolases
|
85(1,1,1,5) |
Details |
|
CCL4
|
84(1,1,1,4) |
Details |
|
p38
|
83(1,1,1,3) |
Details |
|
ST1C1
|
83(1,1,1,3) |
Details |
|
CCL2
|
83(1,1,1,3) |
Details |
|
CCL3
|
83(1,1,1,3) |
Details |
|
ICRF
|
82(1,1,1,2) |
Details |
|
CCR7
|
81(1,1,1,1) |
Details |
|
Microsomal monooxygenase
|
81(1,1,1,1) |
Details |
|
CCR5
|
81(1,1,1,1) |
Details |
|
CYP1
|
81(1,1,1,1) |
Details |
|
CCL27
|
81(1,1,1,1) |
Details |
|
CaV2.3
|
81(1,1,1,1) |
Details |
|
superoxide dismutase
|
114(0,3,5,14) |
Details |
|
aldehyde dehydrogenase (protein family or complex)
|
108(0,3,5,8) |
Details |
|
alcohol dehydrogenase (protein family or complex)
|
74(0,2,4,4) |
Details |
|
VR1
|
74(0,2,4,4) |
Details |
|
intercellular adhesion molecule 1
|
63(0,1,5,13) |
Details |
|
olfactory receptor
|
58(0,1,1,28) |
Details |
|
IL 6
|
57(0,1,3,17) |
Details |
|
catalase
|
51(0,1,2,16) |
Details |
|
CA1
|
50(0,1,4,5) |
Details |
|
COMT
|
49(0,1,3,9) |
Details |
|
calcineurin (protein family or complex)
|
44(0,1,3,4) |
Details |
|
IL 8
|
44(0,1,2,9) |
Details |
|
C10
|
39(0,1,2,4) |
Details |
|
ATPases
|
38(0,1,1,8) |
Details |
|
cyclin
|
38(0,1,2,3) |
Details |
|
beta adrenoceptor (protein family or complex)
|
36(0,1,1,6) |
Details |
|
TRPA1
|
35(0,1,1,5) |
Details |
|
cytochrome c
|
35(0,1,1,5) |
Details |
|
IkappaBalpha
|
34(0,1,1,4) |
Details |
|
enoyl CoA hydratase
|
34(0,1,1,4) |
Details |
|
beta glucosidase
|
33(0,1,1,3) |
Details |
|
alpha adrenoceptor
|
33(0,1,1,3) |
Details |
|
glutathione reductase
|
33(0,1,1,3) |
Details |
|
caspase (protein family or complex)
|
33(0,1,1,3) |
Details |
|
HEMA
|
32(0,1,1,2) |
Details |
|
Bcl xL
|
32(0,1,1,2) |
Details |
|
CYP2E1
|
32(0,1,1,2) |
Details |
|
p65
|
32(0,1,1,2) |
Details |
|
IKKbeta
|
32(0,1,1,2) |
Details |
|
GADD45
|
32(0,1,1,2) |
Details |
|
cyclin D1
|
32(0,1,1,2) |
Details |
|
transferase
|
32(0,1,1,2) |
Details |
|
GABA at
|
31(0,1,1,1) |
Details |
|
p50
|
31(0,1,1,1) |
Details |
|
somatostatin
|
31(0,1,1,1) |
Details |
|
IkappaB kinase
|
31(0,1,1,1) |
Details |
|
CYP1B1
|
31(0,1,1,1) |
Details |
|
beta ketothiolase
|
31(0,1,1,1) |
Details |
|
NF E2
|
31(0,1,1,1) |
Details |
|
Mop
|
31(0,1,1,1) |
Details |
|
MTA 1
|
31(0,1,1,1) |
Details |
|
lipase a
|
31(0,1,1,1) |
Details |
|
odorant binding proteins
|
31(0,1,1,1) |
Details |
|
CD86 antigens
|
27(0,0,2,17) |
Details |
|
lactate dehydrogenase (protein family or complex)
|
27(0,0,3,12) |
Details |
Docunmentation of Network Visuallization
 Network initializing...Please wait
Network initializing...Please wait
Legend:

: Current entry node;

: Target node;

: Compound node;

 Network initializing...Please wait
Network initializing...Please wait  : Current entry node;
: Current entry node;
 : Target node;
: Target node;
 : Compound node;
: Compound node;